Terence Williams Lab Research
The overarching goal of our research is to understand the mechanisms of the inherent and acquired resistance of cancer cells to current available treatment modalities, particularly radiotherapy and chemotherapy at the genetic, transcriptomic, and proteomic levels.
Such knowledge allows us to design and implement novel therapy strategies for poor-prognosis cancers, including gastrointestinal (pancreatic, esophago- gastric, colorectal, and hepatobiliary), thoracic (non-small cell lung cancer), thyroid, and head and neck malignancies. We use various mouse models (transgenic, genetically-engineered, patient-derived xenograft, and orthotopic models) as well as patient tissues (tumor and normal) in combination with basic molecular and cellular techniques to address the fundamental cancer biological questions about how cancers initiate, develop, and progress to aggressive, therapeutically-resistant, and metastatic disease.
We identify and validate our biomarkers in clinically-relevant disease model systems for the diagnosis, treatment and prognosis of cancer patients. We collaborate with many other labs at City of Hope, across the U.S., and internationally in five main overlapping research areas with the goal to drive development of new clinical trials:
- Caveolae-mediated endocytosis and nutrient scavenging
- Oncogene mediated radioresistance
- Identifying and targeting novel DNA repair and DNA damage response pathways modulating tumorigenesis and therapeutic response
- Developmental therapeutics (e.g. protein-drug conjugates, RNA, radiopharmaceuticals, radioimmunotherapy)
- Molecular profiling and predictive biomarkers in immuno-oncology
Caveolae-Mediated Endocytosis And Nutrient Scavenging
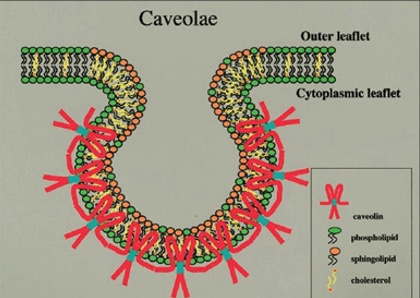
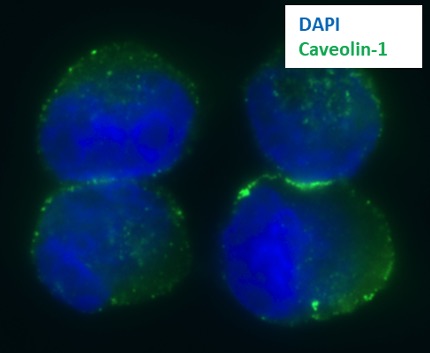
Our laboratory has discovered that caveolae, 50-100 nanometer invaginations of the plasma membrane, can serve as a major pathway for the entry of albumin-bound or -conjugated drugs, as well as nutrient scavenging. We find that caveolin-1 (Cav-1), the principal structural protein of caveolae, is critical for caveolae-mediated endocytosis. We also discovered that Cav-1 is over-expressed and associated with higher tumor grade and worse clinical outcomes in pancreatic cancer and other cancers.
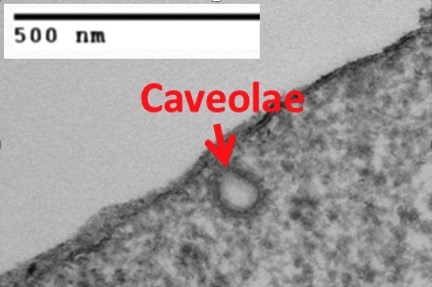

Moreover, Cav-1 promotes the proliferation, migration and resistance of cancer cells to gemcitabine/fluorouracil and radiation, in part through de-regulation of apoptosis and DNA repair. Recently, we have unraveled that high levels of Cav-1 expression lead to increased uptake of albumin-bound chemotherapy drugs (i.e. nab-paclitaxel or Abraxane®) in tumor cells, and that Cav-1 levels are important for nab-paclitaxel response both in vitro and in vivo.
Furthermore, we also find that pretreatment with gemcitabine before nab-paclitaxel increases Cav-1 expression leading to enhanced nab-paclitaxel uptake, and that sequential delivery significantly delays tumor growth both in cells and animal models compared with concurrent delivery.
- Elucidate a role for caveolae-mediated endocytosis in response to novel albumin-conjugated drugs
- Develop novel anti-tumor therapeutics targeting tumors with high levels of caveolae-mediated endocytosis
- Explore the functions of Cav-1 and caveolae mediated endocytosis in nutrient scavenging and tumor metabolism
Oncogene-Mediated Radioresistance
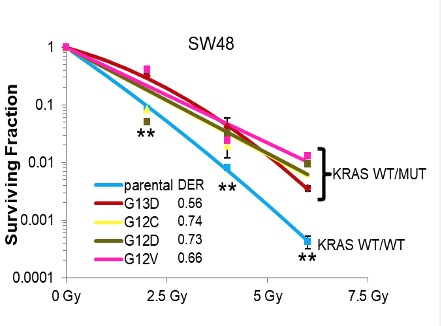
KRAS mutations occur frequently in lung, pancreatic, and colorectal cancer while BRAF mutations are common in thyroid cancer and melanoma. Both are common oncogenic mutations which confers resistance to radiation and genotoxic therapies. Our laboratory research focuses on how KRAS or mutated BRAF leads to heightened DNA repair in the nucleus.
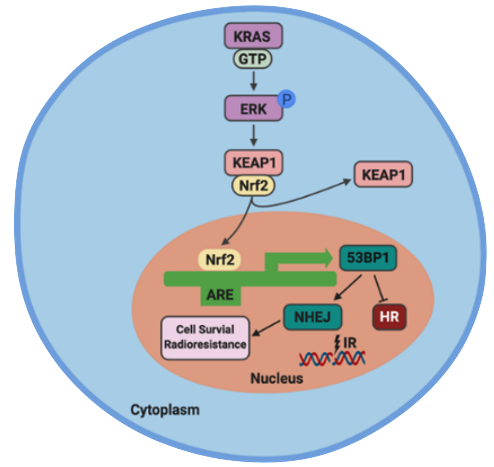
We believe a better understanding of the molecular mechanisms could lead to novel and more targeted therapies in combination with radiation.
Recently, we have found that in colorectal cancer cells, KRAS mutations appears to render radioresistance via enhancing the intracellular KEAP/NRF2 anti-oxidative response, leading to increased transcription of 53BP1. 53BP1 is an important factor controlling DNA repair, including non-homologous end joining (NHEJ).
In a separate study in thyroid cancer and melanoma, we elucidated that BRAF mutation leads to radioresistance by upregulating NHEJ repair factors to promote double-strand break repair after radiation. Excitingly, combining a small molecule inhibitor targeting BRAFV600E mutant potently sensitizes BRAF mutant thyroid cancer cells to radiotherapy.
Our aims are to:
- Understand the role for KRAS and BRAF mutations in the differential modulation of distinct nuclear DNA repair machineries.
- Reveal the functions of KRAS and BRAF mutations in cancer cell resistance to different treatments including immunotherapy and chemotherapy.
- Identify and validate novel strategies targeting KRAS and BRAF mutants in combination with other therapies.
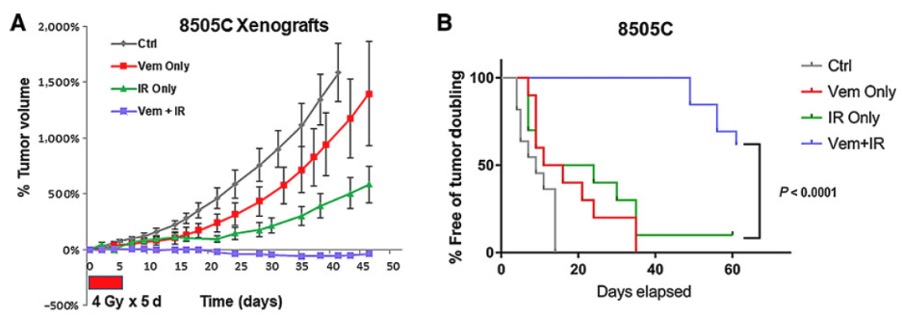
Novel DNA Repair And DNA Damage Response Pathways Modulating Tumorigenesis And Therapeutic Response
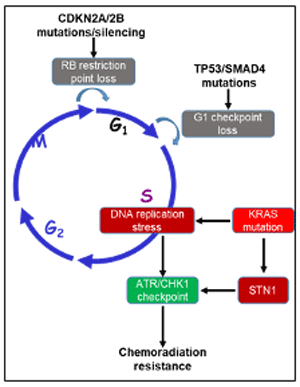
Accurate and complete DNA replication and repair is pivotal for the inheritance of genetic information and maintenance of organisms. Errors in DNA metabolism is a main source of the genome instability of cancer cells. DNA repair and DNA damage response systems surveil the integrity of the genome. However, during the multiple-step process of tumorigenesis and treatment resistance, cancer cells have hijacked these surveillance systems. We find that alterations of the major cell growth signaling pathways such as RAS-MAPK and PI3K-AKT-mTOR signaling promotes distinct DNA repair systems and changes the activity of DNA damage response modulators (e.g. ATM-CHK2 and ATR-CHK1) during late stages of tumorigenesis and treatment relapse. Moreover, uncoupling of the DNA damage response and cell cycle checkpoints leads to uncontrolled growth of tumor cells and targeting the dysregulation of cell cycle checkpoints (e.g. G2/M) can sensitize cancer cells to chemoradiotherapy.
Our aims are to:
- Elucidate the alterations of DNA repair and DNA damage response in cancers with oncogenic mutations in combination with other characteristic cancer gene mutations (e.g. TP53, CDKN2A)
- Identify post-transcriptomic and post-translational mechanisms that mediate altered DNA repair and DNA damage response in cancers with oncogenic mutations
- Develop novel therapeutics targeting oncogenes and aberrant DNA repair to enhance efficacy of therapies
- Understand the non-canonical nuclear functions of signaling molecules (e.g. PTEN, INPP4, mTOR) in DNA damage response and repair
Developmental Therapeutics
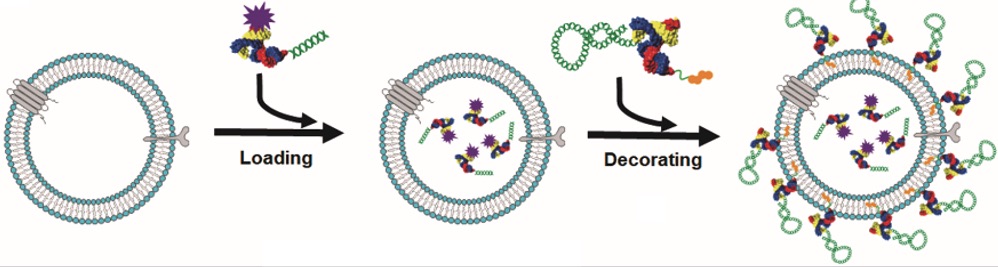
To accelerate the development of new strategies to improve human cancer treatment with improving the quality of life and overall survival time, many macromolecular anti-cancer drugs have been developed based on enhanced permeability and retention (EPR) effect. We are developing RNA-based therapeutics using 3 way junction (3WJ) bacteriophage-derived RNA particles or exosome vesicles (EV) delivered siRNA for important cancer-driving targets such as surviving and KRAS.
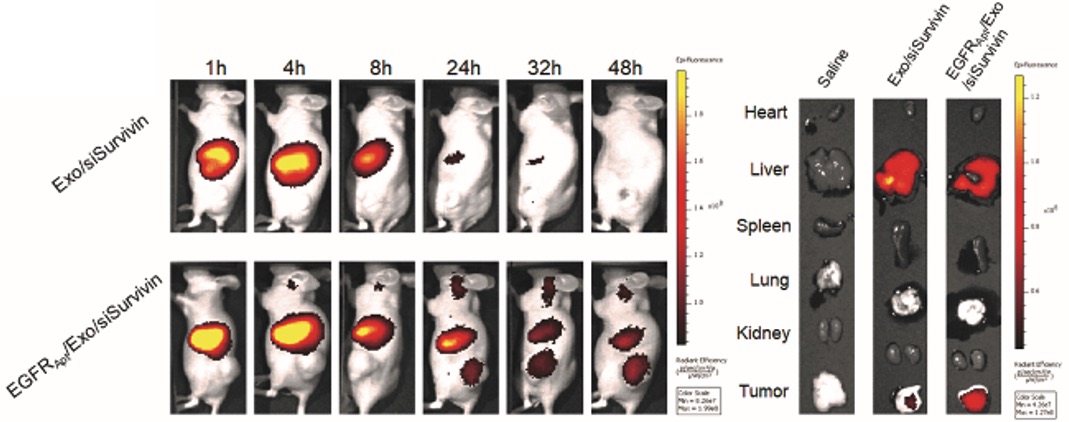
In addition, albumin has been characterized as an ideal drug carrier in the development of novel chemotherapeutic compounds. Recently, our lab has developed novel HSA-chemotherapy conjugates that are potent, effective, safe, and demonstrates improved efficacy in high Cav-1 expressing tumors. We seek to develop and test additional albumin-conjugated therapeutics and develop improved biomarkers of their efficacy. In addition, the ubiquitination system plays a key role in DNA damage response and repair and is deregulated in most cancers. We are identifying specific small molecule inhibitors of E3 ubiquitin ligases important for DNA damage response and repair by using high throughput (content) screening platforms. We are working to develop novel developmental therapeutics, which can then be moved out of preclinical testing into early phase clinical trials.
Our aims are to:
- Develop efficient and targeted macromolecule therapeutics using novel drug and radiotherapy delivery systems (e.g. nanoparticle, exosome, albumin conjugate, antibody-based)
- Identify and validate small molecule inhibitors or genes involved in DNA damage repair and response by high throughput (content) screening or genetic screening approaches (shRNA, CRISPR/Cas9)
- Use high-throughput screening or bioinformatics approaches to identify predictive biomarkers for specific therapies
Molecular Profiling And Predictive Biomarkers
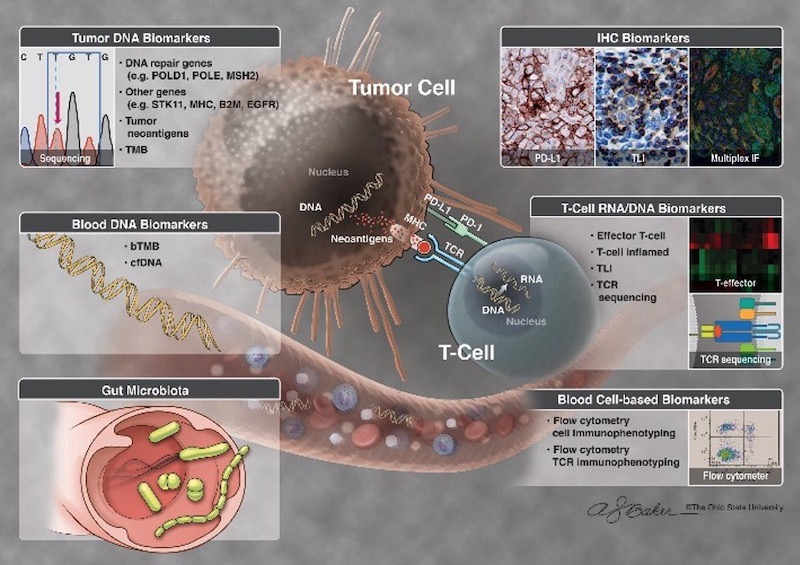
Our lab also focuses on correlative studies from blood and tumor tissues of patients with various cancers to identify prognostic and predictive biomarkers using large-scale or systems-based profiling approaches (e.g. genomic, transcriptomic, proteomic, cellular phenotyping) as well as important new therapeutic targets.
These biomarkers can correlate with treatment efficacy and patient survival to guide personalized cancer care. For example, we have identified miRNA and RNA signatures that are significantly associated with the prognosis of pancreatic, rectal, and lung cancers.
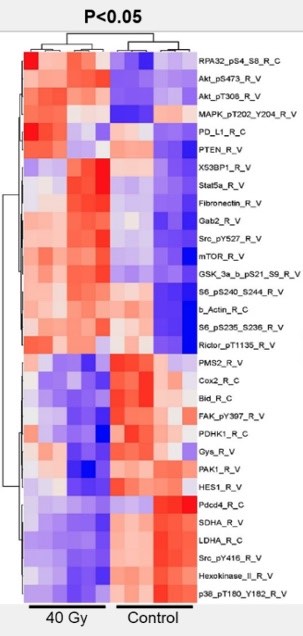
Moreover, using reverse phase protein array of preclinical patient-derived xenograft models, we have found signaling pathways that appear to be mediating chemoradiation in lung cancer. Examples of current projects include: (1) performing RNA profiling in large cohorts of tumors from patients with lung and pancreatic cancers who were treated with different therapies including radiotherapy and chemotherapy; (2) performing profiling of blood immune cell types and other biomarkers to predict response to chemoradiation in combination with immunotherapy from phase II and III trials.
We are seeking to:
- Identify molecular biomarkers that predict for albumin-drug uptake and albumin-based therapeutics.
- Discover novel prognostic molecular biomarkers and predictive molecular biomarkers for radiotherapy, chemotherapy, and immunotherapy in poor-prognosis cancers.
Clinical Trials
City of Hope National Medical Center has hundreds of open clinical trials at any given time, with some of the world’s latest discoveries available to clinical trial patients right here in the greater Los Angeles area. Patients have access to many cancer clinical trials here with access to some of the most advanced, targeted treatments, cellular therapies, advanced radiation technologies, and drugs available.